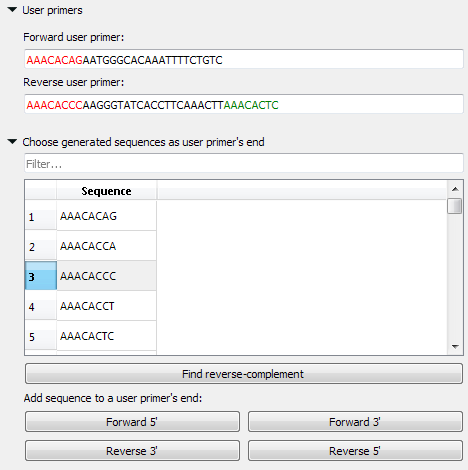
The polymerase chain reaction (PCR) uses a pair of custom primers to direct DNA elongation toward each other at opposite ends of the sequence being amplified. These primers are typically between 18 and 24 bases in length and must code for only the specific upstream and downstream sites of the sequence being amplified. A primer that can bind to multiple regions along the DNA will amplify them all, eliminating the purpose of PCR.
A few criteria must be brought into consideration when designing a pair of PCR primers. Pairs of primers should have similar melting temperatures since annealing during PCR occurs for both strands simultaneously, and this shared melting temperature must not be either too much higher or lower than the reaction's annealing temperature. A primer with a Tm (melting temperature) too much higher than the reaction's annealing temperature may mishybridize and extend at an incorrect location along the DNA sequence. A Tm significantly lower than the annealing temperature may fail to anneal and extend at all.
Additionally, primer sequences need to be chosen to uniquely select for a region of DNA, avoiding the possibility of hybridization to a similar sequence nearby.
Also, there are other parameters to be taken in account - Gibbs free energy, self- and heterodimers availability, etc. All parameters are described below in details.
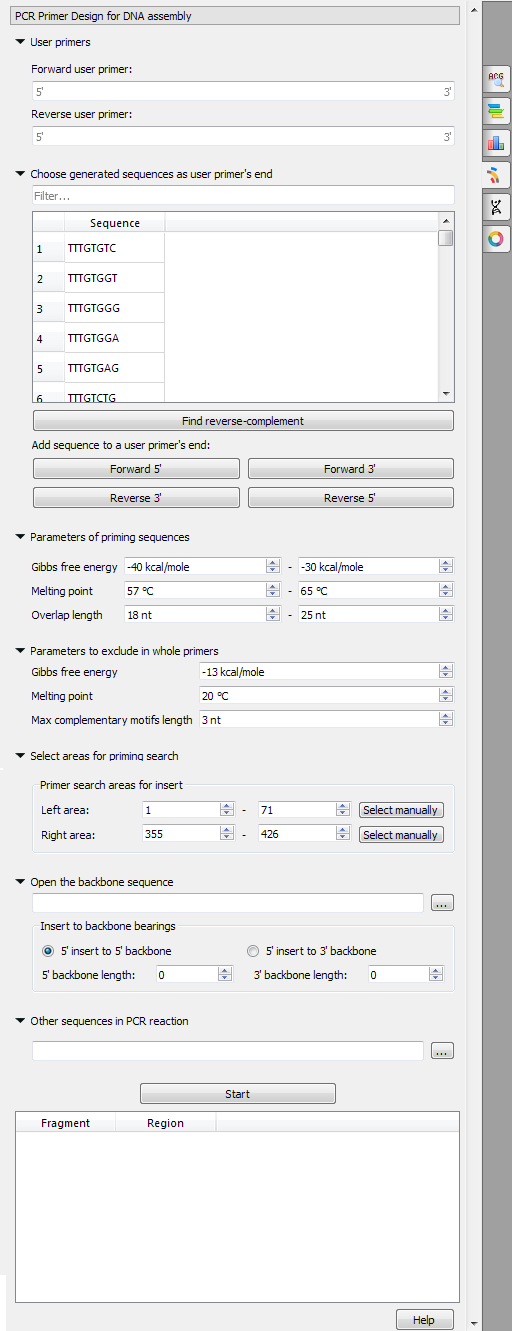
UGENE provides the "PCR Primer Design for DNA Assembly" feature only for nucleic sequences with the "Standard DNA" alphabet. To use it in UGENE open a DNA sequence and go to the "PCR Primer Design for DNA Assembly" tab of the Options Panel:

There are the following parameters:
The designed primers should have the set Gibbs free energy, melting temperature and length. Also, these primers could have hairpins, self- and hetero-dimers, but the mentioned parameters of these found dimers should be limited. Look at the pictures:
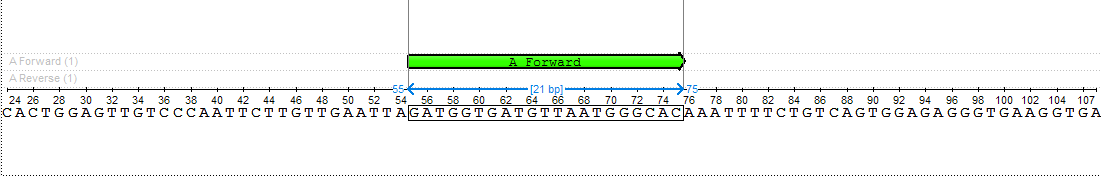
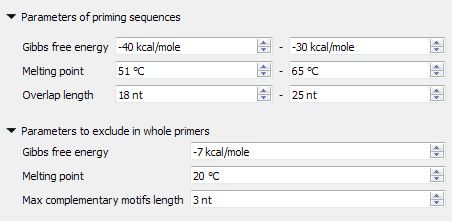
The found primer "GATGGTGATGTTAATGGGCAC" has Gibbs free energy between -40 and -30 kcal/mol, melting temperature between 51 and 65 °C and length between 18 and 25 nucleotides. Also, there are no heirpins, self- and hetero-dimers in this primer, which has Gibbs free energy more then -7 kcal/mol, melting temperature less than 20 °C and base pairs length more than 3 nucleotides.
Forward primers are searched at the left region, reverse primers are searched at the right region. There is an amplified fragment between them: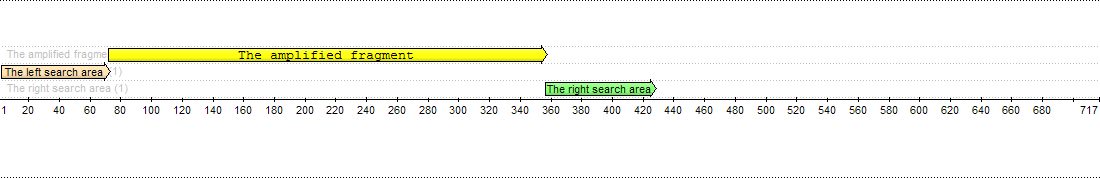
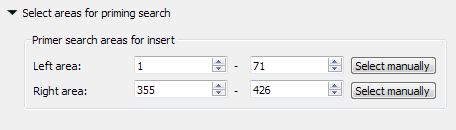
The left search area is from 1 to 71 base, the right search area is from 355 to 426 base. The amplified fragment between them.
Backbone sequence is the sequence that is added to 5' end of the primer for the following assembly of the PCR product with the backbone molecule. This parameter is optional. You may set the backbone sequence, the sequence end you want to add backbone to and the length of the sequence from 5' and 3' ends to check if these sequences have unwanted hairpins, self- and hetero-dimers. On the pictures below you may see the differences between "5' insert to 5' backbone" and "5' insert to 3' backbone" ("insert" in this case is the same as "primer").
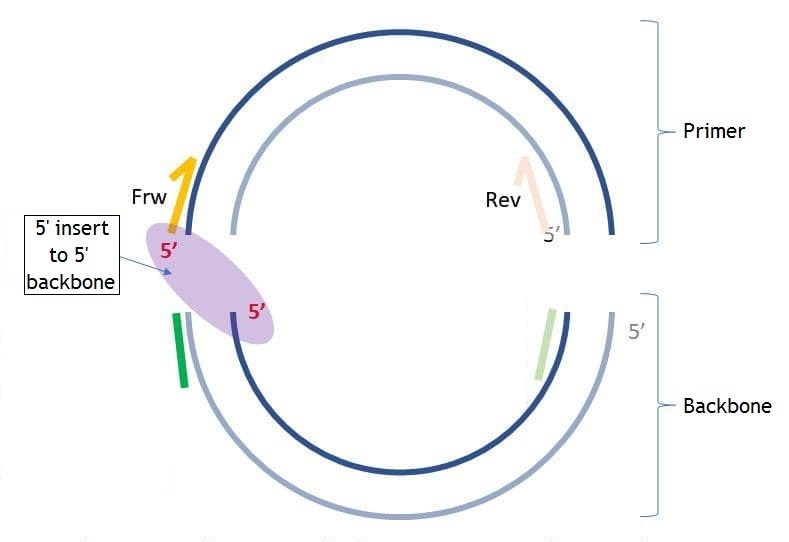
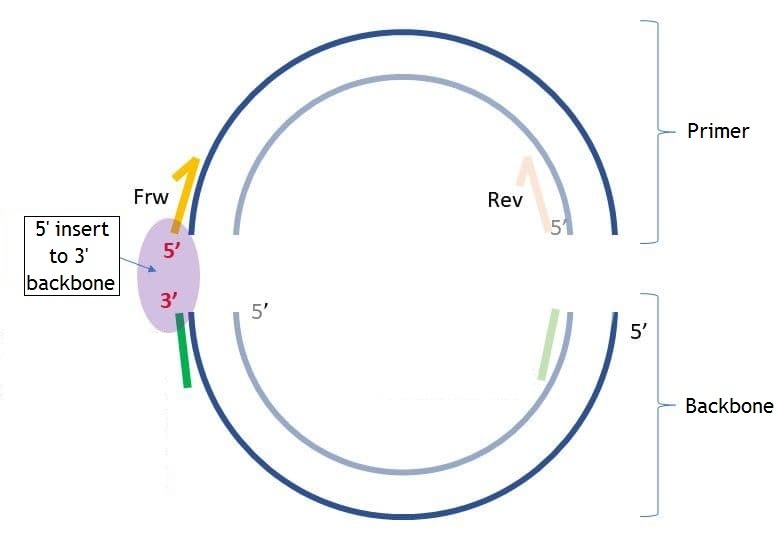
If you set "5' backbone length", then the number of bases you set from 5' end of the backbone sequence will be checked for unwanted connections. The same about "3' backbone length", but from the 3' end. If the checked end has hairpins, self- and homo-dimers you will see the following dialog:
Sequences that are supposed to be in the reaction mixture. Its important primers do not have unwanted connections to these sequences, so it will be checked that there are no hairpins, self- or hetero-dimers between the resulting primer and any sequence from the "Other sequences in PCR reaction" file.
The search result is from 1 to 4 pairs of primers. The results should be as close as possible to the amplified fragment. That means, that the searching process starts from the edge of the amplified fragment to the 5' direction on the direct strand in case of the forward primer and to the 3' direction of the reverse-complementary strand in case of the reverse primer. The results are defined with special names:
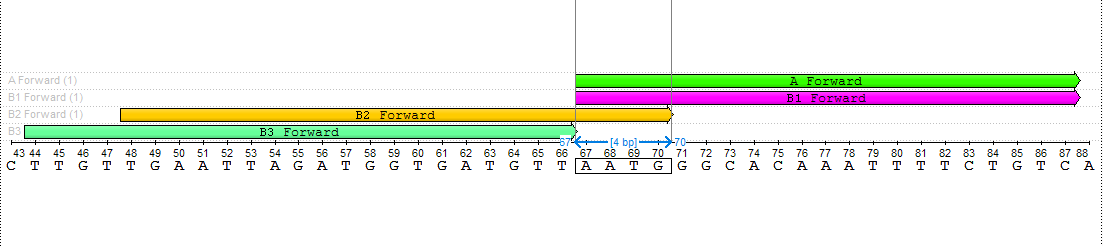
Also, you may see results in the result table. A single click on the result in the result table selects the corresponding annotation, double click - export the resulting primer to the separate file (with the backbone sequence, which will be added during the exporting process).
You may check the validity of the pair of primers by typing this pair into the corresponding fields. Also, you may add your primers 5' or/and 3' 8 bases endings, which do not form simple unwanted connections.