To make a request to a local BLAST database do the following:
- If you’re using BLAST open Tools ‣ BLAST ‣ BLAST Search.
- If you’re using BLAST+ open Open Tools ‣ BLAST ‣ BLAST+ Search.
If there is a sequence opened you can also initiate the request to a local BLAST database from the Sequence View:
- If you’re using BLAST select the Analyze ‣ Query with BLAST item in the context menu or in the Actions main menu.
- If you’re using BLAST+ select the Analyze ‣ Query with BLAST+ item in the context menu or in the Actions main menu.
The Request to local BLAST database dialog will appear:
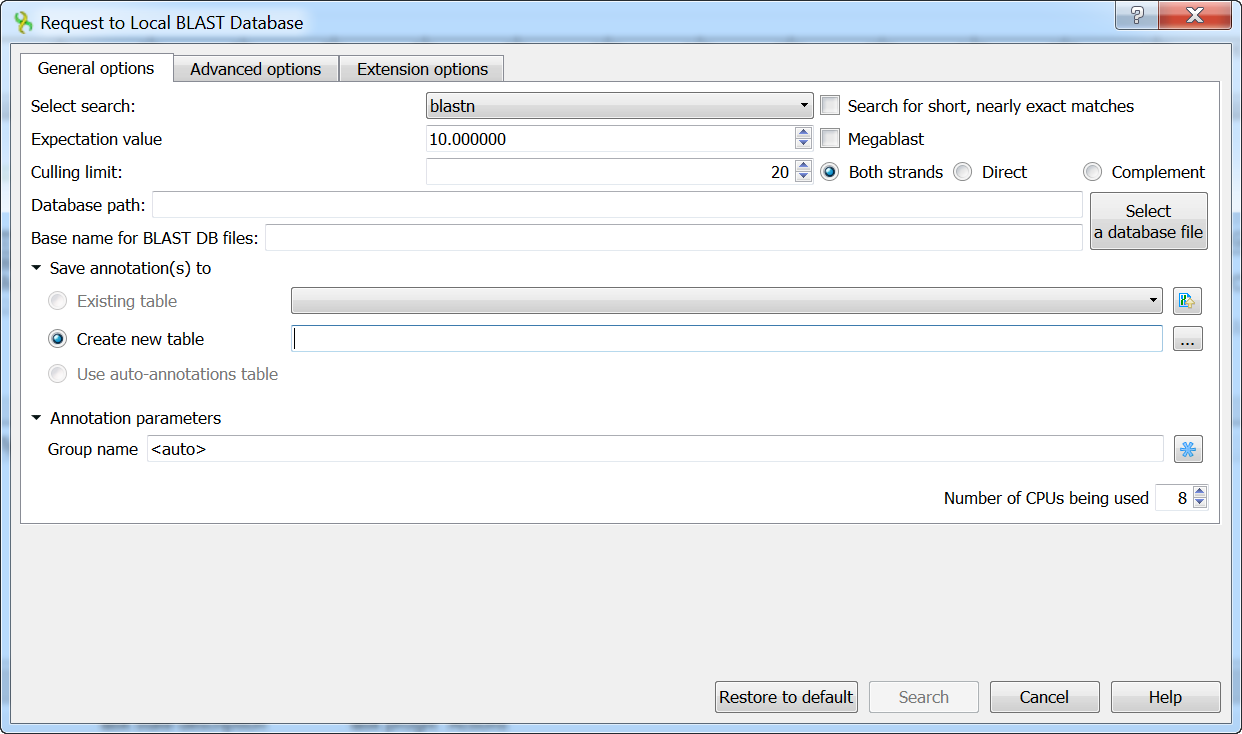
The following general options are available:
Select search - here you should select the tool you would like to use. If the query sequence is a nucleotide sequence then blastn, blastx and tblastx items are available. For a protein sequence the items are blastp and tblastn.
Expectation value - this option specifies the statistical significance threshold for reporting matches against database sequences. Lower expect thresholds are more stringent, leading to fewer chance matches being reported.
Culling limit - the maximum number of hits that will be shown (not equal to number of annotations). The maximum availablle number is 5000.
Search for short, nearly exact matches - automatically adjusts the word size and other parameters to improve results for short queries.
Megablast - select this option to compare query with closely related sequences. It works best if the target percent identity is 95% or more, but it is very fast.
Database path - path to the database files.
Base name for BLAST DB files - base name for the BLAST database files.
You can see the description of the annotation saving parameters here.
The following advanced parameters are available:
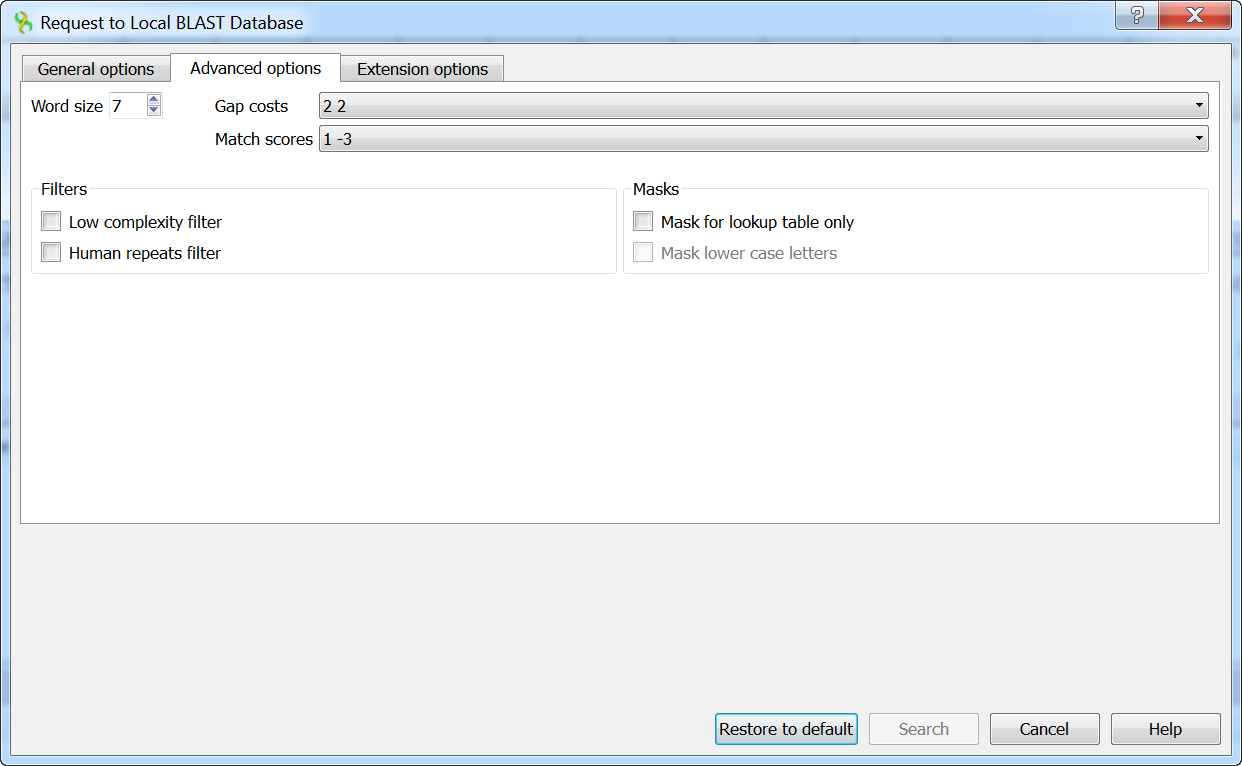
Word size - the size of the subsequence parameter for the initiated search.
Gap costs - costs to create and extend a gap in an alignment. Increasing the Gap costs will result in alignments which decrease the number of Gaps introduced.
Match scores - reward and penalty for matching and mismatching bases.
Filters - filters for regions of low compositional complexity and repeat elements of the human’s genome.
Masks for lookup table only — this option masks only for purposes of constructing the lookup table used by BLAST so that no hits are found based upon low-complexity sequence or repeats (if repeat filter is checked).
Mask lower case letters — with this option selected you can cut and paste a FASTA sequence in upper case characters and denote areas you would like filtered with lower case.
The following extension options are available:
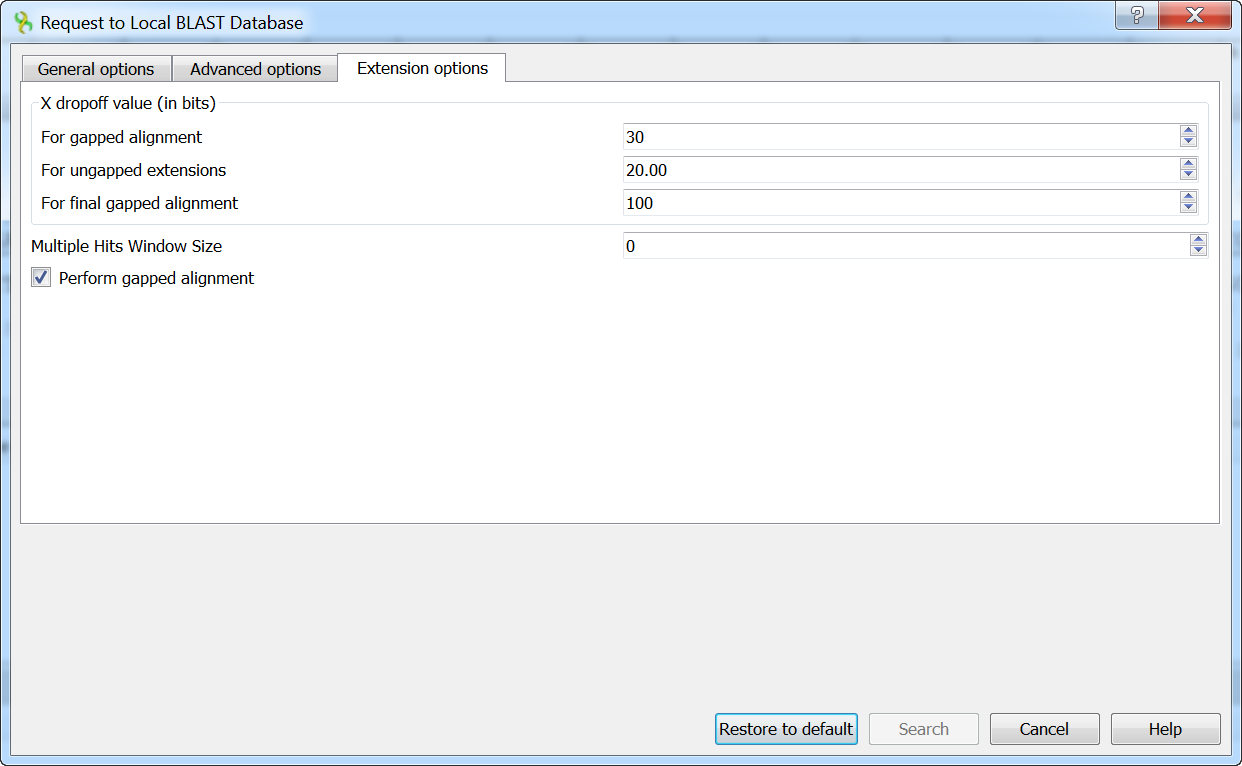
For gapped alignment - X dropoff value (in bits) for gapped alignment.
For ungapped alignment - X dropoff value (in bits) for ungapped alignment.
For final gapped alignment - X dropoff value (in bits) for final gapped alignment.
Multiple hits window size - multiple hits window size.
Perform gapped alignment - performs gapped alignment.