Enabling the automatic annotations highlighting allows you to automatically calculate and highlight annotations on each nucleotide sequence opened.
Currently, the following annotations types support the automatic highlighting:
- Open reading frames
- Restriction sites
- Plasmid features
The corresponding groups of annotations found are stored in the Auto-annotations object in the Annotations editor, for example:

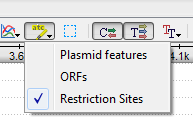
To disable/enable the automatic annotations calculations to use the Automatic annotations highlighting menu button on the Sequence View toolbar:
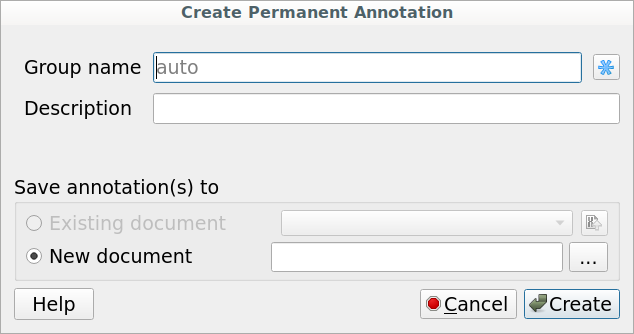
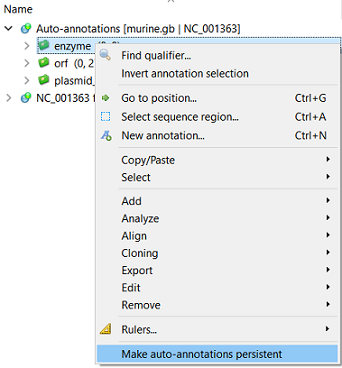
To create a permanent annotation click on the Make auto-annotations persistent context menu item and choose the annotation parameters in the Create Permanent Annotation dialog.
The following dialog will appear: