To search for a pattern(s) in a sequence go to the Search in Sequence tab of the Options Panel in the Sequence View.
Input the value you want to search in the text field. To search multiple patterns input the patterns separated by a new line in the pattern text field. To add a new line symbol Ctrl+Enter may be used. You can input the value as a sequence or name of the sequence in the FASTA format and sequence after that. Press the Next button to search in the direction “From left to right”. Press the Previous button to search in the direction “From right to left”. If the pattern is found, the result will be focused and highlighted in the Sequence area. You can continue the search in any direction from this position.
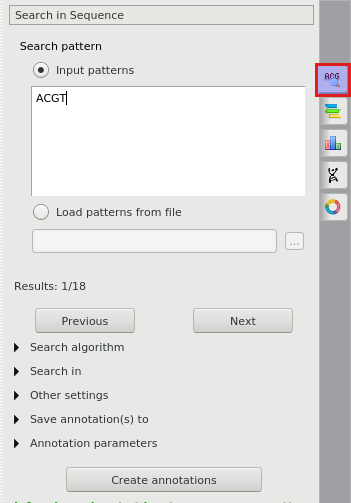
By default, misc_feature annotations are created for regions that exactly match the pattern. Find below the description of the available settings.
Load Patterns from File
Use this option to load patterns from a file. When this option is active the Search for field is disabled.
Search Algorithm
This group specifies the algorithm that should be used to search for a pattern. The algorithm can be one of the following:
- InsDel — there could be insertions and/or deletions, i.e. a pattern and the searched region can vary in their length. You can specify the percentage of the pattern and a searched region match in the field nearby. Note that this value also depends on the pattern length and is disabled when the pattern hasn’t been specified.
- Substitute — a pattern may contain characters different from the characters in the searched region. When this algorithm has been selected you can also specify the match percentage and additionally it is possible to take into account ambiguous bases.
Regular expression — a regular expression may be specified instead of a pattern. For example character ‘.’ matches any character, ‘.*’ matches zero or more of any characters. There is also the Result no longer than option that specifies the maximum length of a result.
- Exact - find a place where one or several patterns are found within a larger pattern.
Search in
In this group you can specify where to search for a pattern: in what region and in which strand (for nucleotide sequences). Also for nucleotide sequences, it is possible to search for a pattern on the sequence translations.
Strand — for nucleotide sequences only. Specifies on which strand to search for a pattern: Direct, Reverse-complementary or Both strands.
Search in — for nucleotide sequences you can select the Translation value for this option. In this case, the input pattern will be searched in the amino acid translations.
Region — specifies the sequence range where to search for a pattern. You can search in the whole sequence, specify a custom region or search in the selected region.
Other Settings
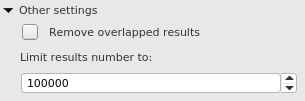
This group contains additional common settings:
Remove overlapped results — annotates only one of the overlapped results.
Limit results number to — limits the number of the searched results to the specified value.
Annotations Settings
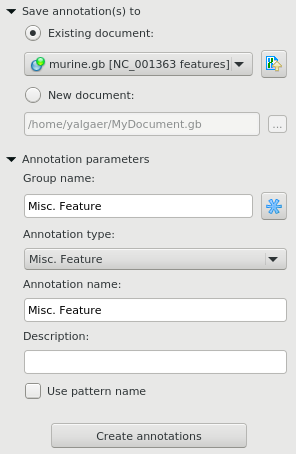
In the Save annotation(s) to group, you can set up a file to store annotations. It could be either an existing annotation table object or a new annotation table.
In the Annotation parameters group, you can specify the name of the group and the name of the annotation. If the group name is set to <auto> UGENE will use the group name as the name for the group. You can use the ‘/’ characters in this field as a group name separator to create subgroups. If the annotation name is set to by type UGENE will use the annotation type from the Annotation type: table as the name for the annotation. Also you can add a description in the corresponding text field. To use a pattern name for the annotations check the corresponding checkbox.
After that click the Create annotations button. The annotations will be created.
Searching for one or several patterns and names of the result annotations
If you search for one pattern only, then input the required name into the Annotation name field and leave the Use pattern name checkbox unchecked.
You can also search for several patterns at a time by:
Inputting several patterns into the search field (click <Ctrl> + <Enter> keys to insert to a new line):
- Inputting several patterns into the search filed in FASTA format:
- Loading patterns from a FASTA file
Even when you search for several patterns, the names of the found annotations will be identical by default (the name is specified in the Annotation name field).
If you want to assign different names to annotations found for different patterns, then you should:
- Input the patterns in FASTA format (the latter two cases above)
- Check the Use pattern name checkbox in the Annotation parameters group
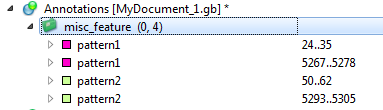
Here is an example of the found annotations in the Annotations Editor: