...
- User primers - the pair of forward and reverse primers user can check on the absence of unwanted connections (like hairpins, self- and hetero-dimers).
- Choose generated sequences as user primer's end - user primers could be extended with the 8-bases sequences on 5' and 3' ends. These sequences do not form too simple unwanted connections (for example, "AAAATTTT", obviously, has a hairpin, so there is no such a sequence in the list).
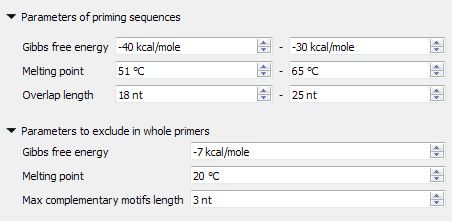
- Parameters of priming sequences - the range of melting temperature, Gibbs free energy and length primers SHOULD have.
- Parameters to exclude in whole primers - the minimum melting temperature, the maximum Gibbs free energy and the maximum base pairs length which may be present in the self- and hetero-dimers in the considered primer.
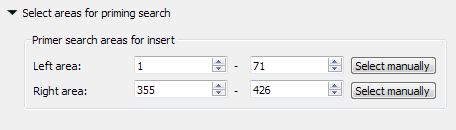
- Select areas for priming search - select areas to search forward and reverse primers in.
- Open the backbone sequence - choose a sequence to be added to 5' or 3' end of the result primer.
- Other sequences in PCR reaction - the list of sequence to check the unwanted connections with. The unwanted connections should fit to "Parameters to exclude in whole primers".
Designing primers
Physical quantity
The designed primers should have the set Gibbs free energy, melting temperature and length. Also, these primers could have hairpins, self- and hetero-dimers, but the mentioned parameters of these found dimers should be limited. Look at the pictures:
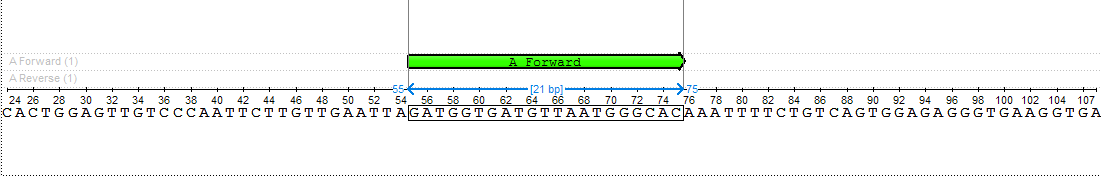
The found primer "GATGGTGATGTTAATGGGCAC" has Gibbs free energy between -40 and -30 kcal/mol, melting temperature between 51 and 65 °C and length between 18 and 25 nucleotides. Also, there are no heirpins, self- and hetero-dimers in this primer, which has Gibbs free energy more then -7 kcal/mol, melting temperature less than 20 °C and base pairs length more than 3 nucleotides.asfsd
Regions
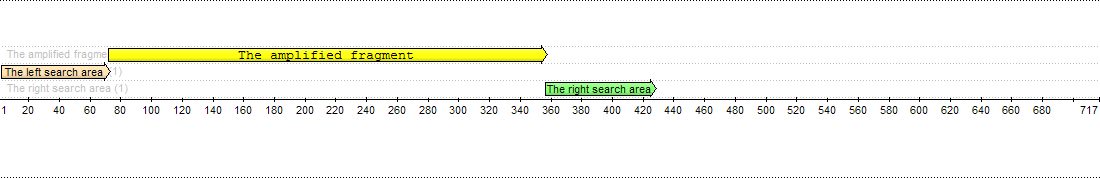
Forward primers are searched at the left region, reverse primers are searched at the right region. There is an amplified fragment between them:
The left search area is from 1 to 71 base, the right search area is from 355 to 426 base. The amplified fragment between them.
Backbone sequence
Backbone sequence is the sequence that is added to 5' end of the primer for the following assembly of the PCR product with the backbone molecule. This parameter is optional. You may set the backbone sequence, the sequence end you want to add backbone to and the length of the sequence from 5' and 3' ends to check if these sequences have unwanted hairpins, self- and hetero-dimers.
Type two primers for running In Silico PCR. If the primers pair is invalid for running the PCR process then the warning is shown. Also, primers for the running In silico PCR can be chosen from a primer library. Click the following button to choose a primer from the primers library:
...