The Primer3 plugin is a port of the Primer3 tool, v. 2.6.1. It is intended to pick primers from a DNA sequence.
Usage
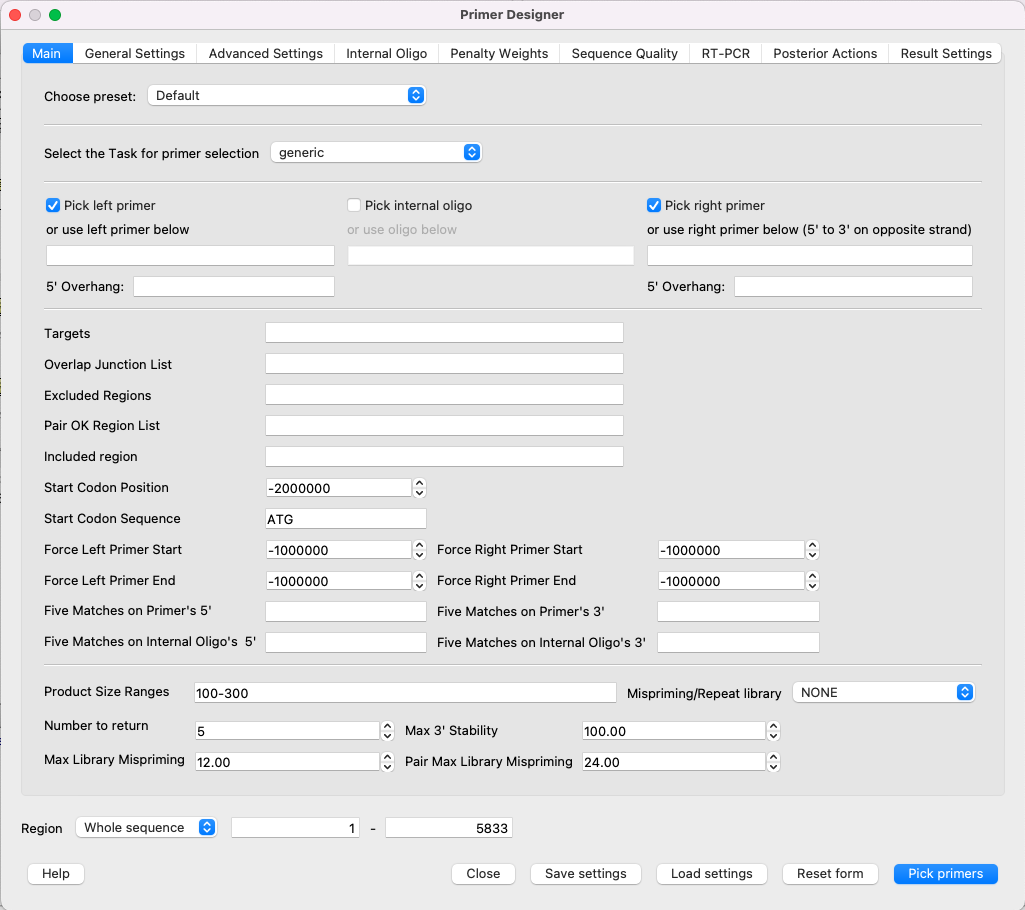
To use the Primer3, open a DNA sequence and select the Analyze ‣ Primer3 context menu item. The dialog will appear:
All available parameters are the same as the original Primer3 has.
NOTE: The "Primer Designer" dialog has a detailed description of each parameter - just move the mouse cursor to the parameter you're interested about (or its name) and the tooltip appears. Due to this fact, there is no detailed description of each parameter in this file.
Report
Then calculation is finished, the notification appeared in the bottom right corner. Click on it to open the report file.
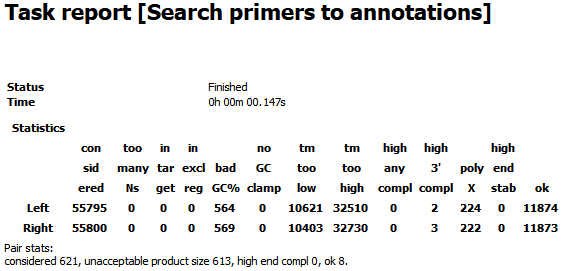
The report looks as follow and contains the information about how many primers were considered during the searching process and how many of them were filtered due to one or another reason.
Results
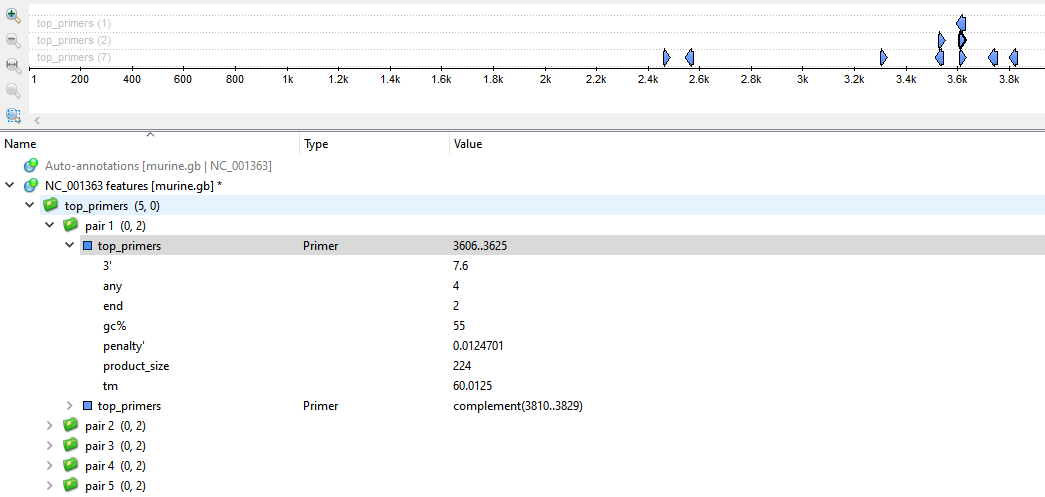
All found primers will be available as annotations and marked on the target sequence:
The result annotations could have a set of the following qualifiers:
- 3' - free Gibbs energy of 5 bases on 3' end.
- any - value, which shows tendency of this primer to bind it's reverse-complement chain. This value calculates as follows: primer aligns on its reverse-complement representation, each match counts +1 point, each mismatch -1 point, each gap -2 points, each N -0.25 points.
- end - the same as any, but only about 5 bases on 3' end.
- gc - GC-content.
- penalty - penalty weight (see this article for details).
- hairpin - a hairpin loop melting temperature (if this kind of loop exists in the primer).
- product_size - size of the product.
- tm - primer melting temperature.
However there is one additional feature available which is not originally a part of Primer3 tool. It allows user design primers for RT-PCR experiments by choosing which exons/introns to span with the primer product. This feature is described in detailed below. When you select the parameters you can save and load settings with a help of the corresponding buttons in the right corner of the dialog.